• Animal molecular phylogenetics and DNA barcoding of Mollusca and Pancrustacea
Phylogeny is the hard core of all zoological study and each zoological study adds to phylogeny. This research line is, in particular, devoted to the molecular characterization of both mitochondrial and nuclear markers (rDNA, microsatellites, inter-SINEs, single copy genes), but also whole (mito-)genome sequencing. Together with morphological analyses, we aim at the systematics that only derives from the most plausible natural phylogeny of the groups under study.
Recently, the DNA barcoding approach has been added to the study of some lineages (Pancrustacea, Phasmida).
Mollusca
Mollusca have always been and are still considered one of the biggest question marks across the evolutionary tree of Metazoa. In recent times, a close relationship with Annelida has been strongly advocated, if not accepted (e.g., Struck and Fisse, Mol. Biol. Evol. 2008 25: 728-736; Andrade et al., Mol. Biol. Evol. 2014 31:3206-3215). Conversely, the internal relationships of molluscs are widely debated: almost all conceivable combinations of molluscan classes have been proposed as monophyletic clades (Sigwart and Lindberg, Syst. Biol. 2015 64:384-395).
Unravelling the phylogeny of single molluscan classes is therefore mandatory to gain a clearer understanding of the evolution of the phylum as a whole. A research line of our lab focuses on the phylogeny of the class Bivalvia, with special emphasis on deep nodes.
To this purpose, mitochondrial markers were used, as well as complete mitochondrial genomes. In this regard, the sequencing of the complete mitochondrial genome of Solemya velum in 2013 (Plazzi et al., BMC Genomics 2013 14:409) was a key achievement of our group, in that this species retains many ancestral mitogenomic (as well as morphological) features.
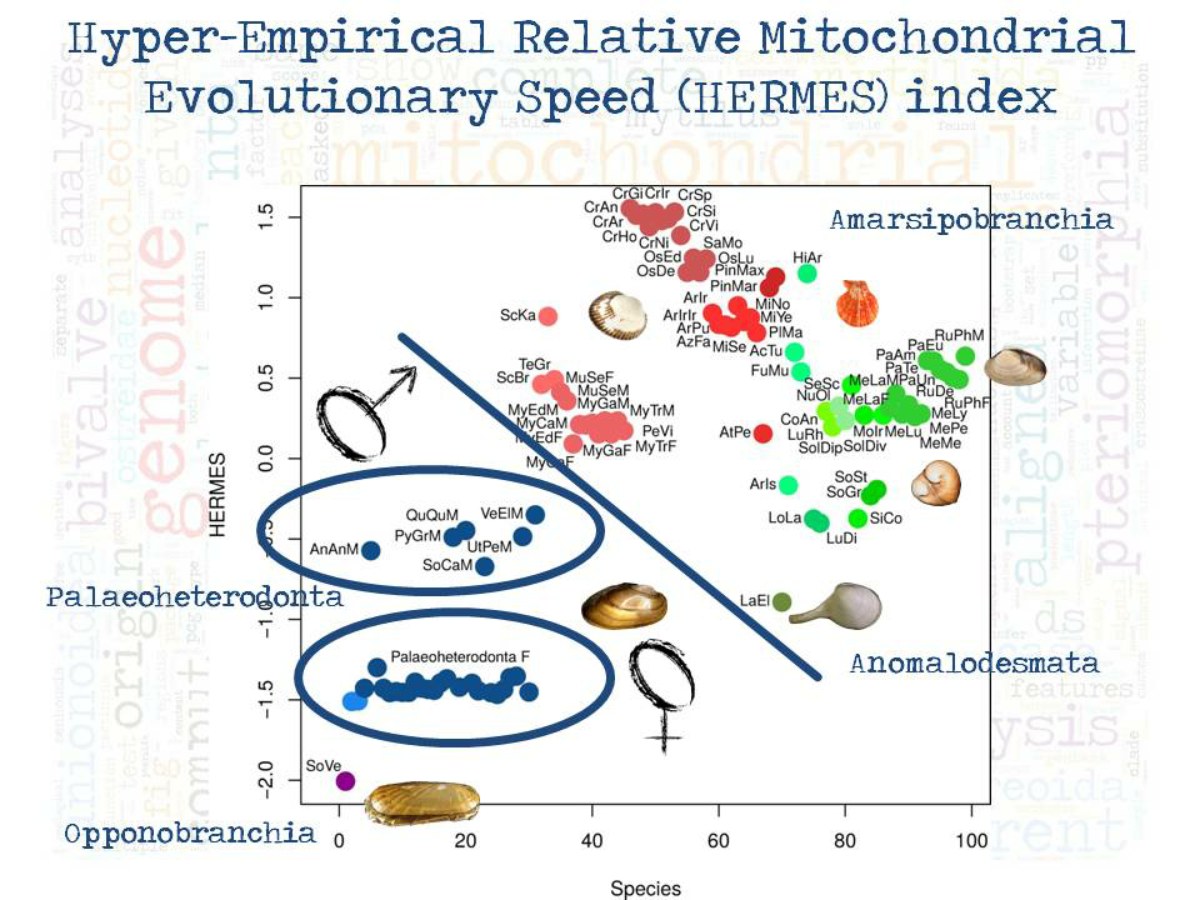
A long-debated issue on bivalve phylogeny is the sister group of the subclass Heterodonta (Venus clams, cockles, quahogs, razor clams, and their kin): the use of mitochondrial markers leads us to favour the Heterodonta+Pteriomorphia hypothesis (where Pteriomorphia includes mussels, oysters, scallops, and their kin), a clade we called Amarsipobranchia (Plazzi et al., PLoS One 2011 6:e27147). Recent findings based on high-throughput data are currently challenging this view, while the fossil record would strengthen our hypothesis (Plazzi et al., Genome Biol. Evol. 2016 8:2544-2564).
Moreover, mitochondrial genomics of bivalves is interesting per se, in that bivalve mitochondrial genomes show many uncommon aspects. Outstanding is the presence in some species of the DUI mechanism for mitochondrial heredity (which is the focus of another research line of our lab), but there are many other peculiarities of bivalve mitochondrial genomes. A sound phylogenetic framework is mandatory to fully evaluate the evolution of these unique organellar genomes.
Pancrustacea
Taxonomy, systematics and phylogenetics currently integrate with molecular markers and, in the last decades, further implemented through the use of molecular markers. We therefore study the standing biodiversity, starting from the morphological approach and adding data from (whole) mitochondrial and nuclear genomes. Taxa under study are represented by the termite genera Reticulitermes and Kalotermes (Hexapoda, Insecta, Isoptera) and the tadpole shrimps genera Triops and Lepidurus (Crustacea, Branchipoda, Notostraca). Data gathered in these projects are a strong support in comparative studies of other zoological aspects, giving a clear-cut picture of the taxonomic/phylogenetic milieu.
Recently, with stick-insects, we entered in the DNA barcoding world. The “Barcode of Life” is a broad project on planetary scale aiming to develop a DNA sequence-based global standard for species determination. For this purpose, the first 600 bp of the cox1 mitochondrial gene have been identified as the more reliable barcoding sequence for almost all animal species (http://www.barcoding.si.edu).
The order Phasmida includes the so-called stick and leaf insects, well known for being one of the clearest examples of cryptic mimetism and for showing a wide variety of reproductive strategies, including parthenogenesis, hybridogenesis and androgenesis (Scali et al., 2003. Biol J Linn Soc 79: 137–150; Schwander et al., 2011. Curr Biol 21:1129-1134; Scavariello et al., 2017. Scientific Reports, 7:41946). The order presently contains more than 3200 species distributed worldwide (except Arctica and Antarctica), with greater abundance in the tropics and sub-tropics. This huge biodiversity calls for a validation of currently recognized taxa and of their phylogeny as, over 3200 valid species, still more than 4800 taxonomic names have been issued. Moreover, beside to the DNA barcoding, an order-scale molecular phylogeny will offer the opportunity to investigate about the origin of phasmids and about ecological and historical factors that shaped their current distribution. We anticipate exciting discoveries during the progress of the campaign, thanks to a huge international collaboration involving researchers/amateurs from all over the world.
Collaborations:
- Oskar Conle (taxonomist, Germany). Owns the world's largest collection, which will be one of the major sources for DNA-samples.
- Frank Hennemann (taxonomist, Germany). Owns the world second largest private collection, which will be one of the major sources for DNA-samples.
- Paul Brock (Phasmid Study Group , UK): a world-wide recognized expert (see: http://phasmida.speciesfile.org/HomePage/Phasmida/HomePage.aspx).
- Pablo Valero and Prof. Dr. Antonio Ortiz (University of Murcia, Spain).
- Prof. Dr. Jane Margaret Costa von Sydow (Instituto Oswaldo Cruz/ Fiocruz, Rio de Janeiro, Brazil).
- Dr. Bruno Kneubühler (Switzerland). World's leading expert in phasmid breeding.
- Joachim Bresseel & Jerome Constant (taxonomists, Institut Royal des Sciences Naturelles, Bruxelles, Belgium).
- Yeisson Gutierrez (University of Münster, Germany).
Links: